AI for Digital Pathology
Key Aspects
With healthcare software engineering experience since 2005 and ISO 13485 certification, ScienceSoft helps clinical and research-focused digital pathology product companies engineer AI features for existing platforms and build AI-powered digital pathology software from scratch.
AI for Digital Pathology: Essence
Digital pathology software with AI uses machine learning and computer vision to process whole-slide images (WSI) and identify, measure, and classify tissue patterns. In clinical settings, it helps pathologists review cases faster, quantify biomarkers more consistently, and prioritize high-risk slides. In research, it helps turn slide data into structured insights for cohort analysis, biomarker exploration, and multimodal studies.
AI In Pathology Market
The global AI in pathology market size is projected to reach USD 1.15 billion by 2033, growing at a CAGR of 27.18%. The key drivers of AI’s growth in digital pathology are the rising number of cancer and chronic disease cases, which create more slide-review work for labs, the need to process slides faster and boost productivity, the ongoing digitization of pathology workflows, and AI becoming more advanced and affordable to deploy.
How AI for Digital Pathology Works
Common use cases
AI for clinical pathology workflows
Triage and case prioritization
AI assigns a risk or priority score to each slide and pushes the most suspicious cases to the top of the worklist, so pathologists review higher-risk or likely positive cases first.
Cancer detection support
AI highlights suspicious regions on pathology slides (heatmaps or overlays), including possible metastases in lymph nodes, to guide the reviewer’s attention and reduce time spent searching.
Cancer grading and clinically important feature detection
AI detects patterns linked to severity (e.g., grading-related features) and provides structured suggestions for the pathologist’s confirmation.
Tumor segmentation and quantitative measurements
AI outlines tumor regions and calculates metrics, such as tumor area percentage, to make measurements faster and more consistent.
Biomarker and immunohistochemistry (IHC) quantification
AI counts stained cells and estimates biomarker scores (e.g., Ki-67) to speed up, standardize, and reduce the labor intensity of routine pathology scoring.
Slide and scan quality control
AI detects focus and artifact issues early (folds, bubbles, staining problems) so slides can be rescanned or routed before they affect review results or AI accuracy.
Second-read support
AI provides a secondary review layer by flagging potentially missed regions or discrepant findings, helping pathologists reduce oversight risk.
AI for research and discovery
Biomarker exploration and validation
AI helps identify image-based features associated with biomarkers, treatment response, or disease progression, supporting biomarker hypothesis generation and validation.
Cell and tissue quantification
AI classifies tissue types and counts specific cell populations across large slide sets, turning images into structured data for quantitative phenotyping, research, and trials.
Tumor microenvironment analysis
AI identifies and quantifies immune cells and spatial relationships between tumors and surrounding tissue to support biomarker research and precision oncology.
Cohort discovery and stratification
AI groups slides or patients by shared morphological patterns, biomarkers, or risk signals, helping research teams build cohorts for retrospective studies, translational research, and clinical trials.
Multimodal analysis
AI links pathology image features with genomics, outcomes, or trial data to uncover stronger disease signatures and support research-grade model development.
Reference architecture for an AI-powered digital pathology platform
Below, ScienceSoft’s solution architects show a reference architecture for a clinical or lab digital pathology product with AI, including how the core pathology workflow can be implemented and how the AI component can be continuously monitored, evaluated, and updated. For research-focused solutions, the architecture would have a similar overarching logic but require a dedicated lane for data de-identification and governance, feature extraction storage, cohort building and search, multimodal data joins with genomics, outcomes, or trial data, and an experiment workspace or research API.
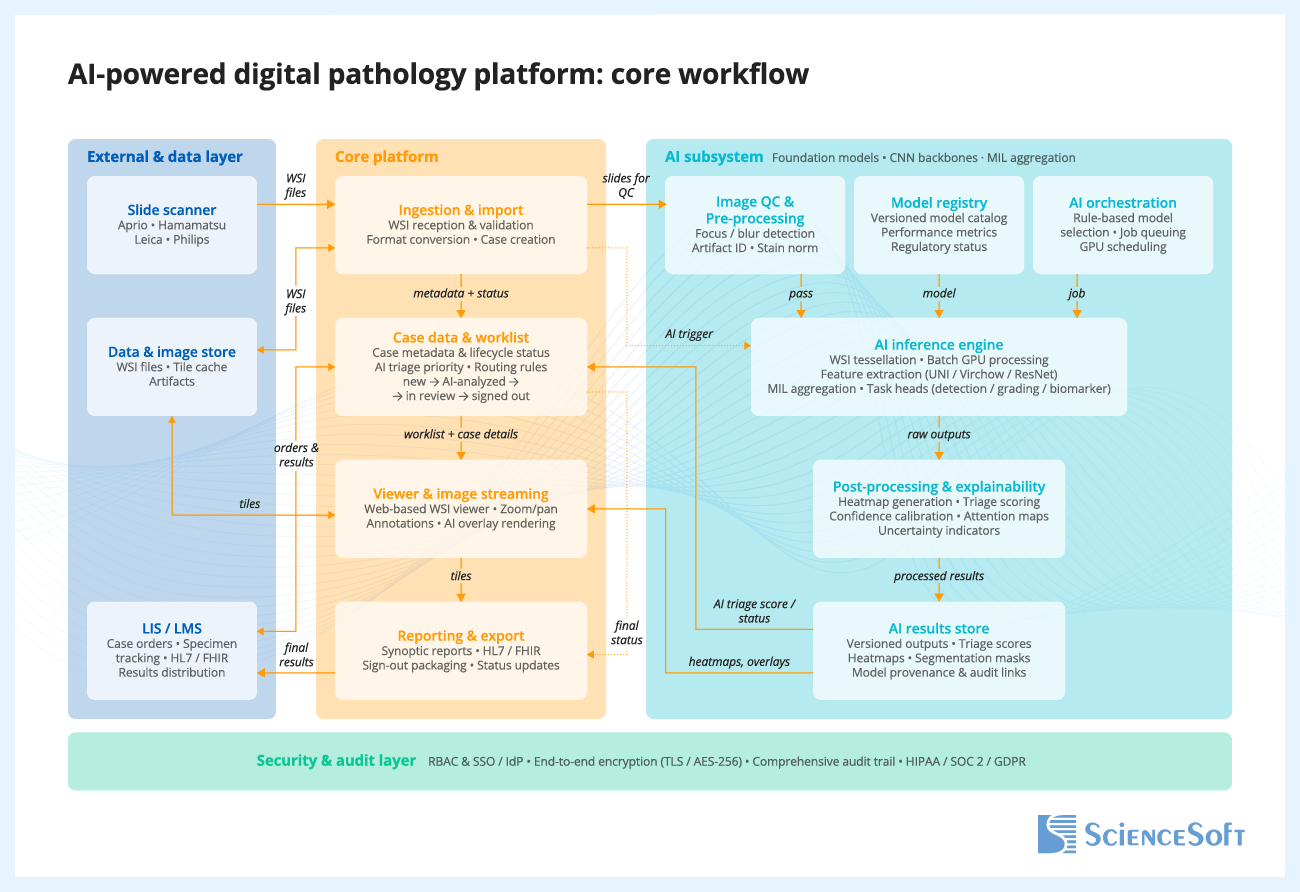
When a scanner creates a whole-slide image (WSI) and sends it to the platform, the platform validates the file, stores it, creates or updates the case record, and routes the slide for AI analysis, if configured. Inside the AI subsystem, the slide goes through image quality checks, model selection, inference, and post-processing, which converts raw model outputs into usable results such as classifications, heatmaps, scores, and measurements. These results are stored with versioning and traceability for the model, inputs, and outputs. The case worklist and viewer are then updated so AI results and overlays are available during review, and prioritization signals can be reflected in the workflow where appropriate. The pathologist opens the case, reviews the slide with or without AI support, and renders the final interpretation. After sign-out, the platform generates the report, updates the case status, and sends results to connected systems such as the LIS or LIMS.
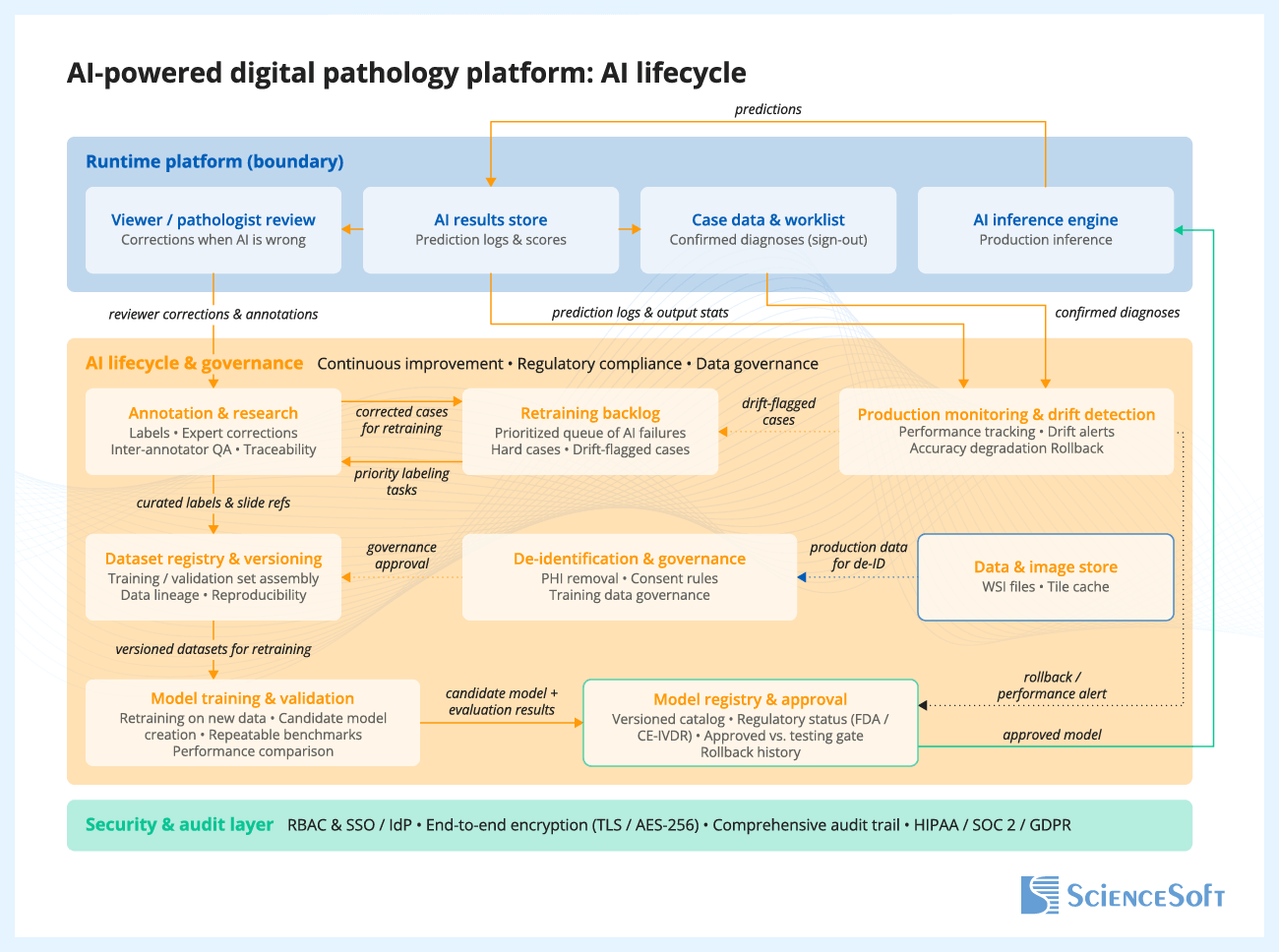
Reviewer feedback on AI outputs is captured and stored for later analysis. The product team reviews this feedback, identifies hard cases and failure patterns, and selects relevant cases for dataset updates. After curation and annotation, the updated data is added to versioned training and evaluation datasets. The model is then retrained and tested against predefined acceptance criteria. Approved model versions are packaged and released for deployment, while production monitoring tracks performance, drift, and operational issues over time.
The Security and Audit Layer operates as a cross-cutting control that spans the entire platform. It verifies every user through secure sign-in, controls what each role is allowed to access or do, and encrypts data both in transit and at rest. It also keeps an immutable audit trail of all important actions, including slide access, AI runs, outputs, and final pathologist decisions. This layer supports compliance with healthcare and data protection requirements such as HIPAA, SOC 2, and GDPR.
Keep AI as a separate back-end component instead of building it directly into the slide viewer or case management system. This one decision solves three problems at once: you can update models without touching the pathologist workflow, run old and new model versions side by side for validation, and swap your entire inference back end (local GPU today, cloud tomorrow) without affecting users.
More importantly, before spending months on the inference pipeline itself, invest two weeks into a structured correction capture mechanism in your viewer. Every time users correct, override, or refine AI output, that disagreement should be stored with the slide reference, the model version, and the original prediction, because those corrections are your most valuable asset for making the next model version better. Teams that skip this step end up with a static model that quietly degrades as scanners change and case mix shifts, with no data even to detect that it is happening.
Technologies We Use to Build AI Products for Digital Pathology
Pathology AI / model-development tech
Pathology foundation models
Slide-level analysis and aggregation
Segmentation and cell analysis
Computer vision backbones
AI frameworks and pathology imaging libraries

MLOps platforms and AI services

Best Practices for Developing AI Products for Digital Pathology
Define a narrow, lower-risk use case before scaling the model
Instead of starting with a broad, high-risk goal such as full cancer grading or complex biomarker scoring, it is usually safer to begin with a narrower workflow, such as scan QC, tissue or region-of-interest (ROI) detection, or second-read triage. These use cases are easier to validate, fit more naturally into real workflows, and help product teams build evidence and user confidence. It is also important to define the intended use and regulatory posture early, for example, whether the AI is meant for research support, review support, or a more advanced diagnostic use case, because that decision shapes the data strategy, validation burden, workflow design, and product claims.
Add AI to existing digital pathology software as a modular workflow layer
Many digital pathology products already have a viewer, storage layer, and core case workflow in place, but lack the controls needed to introduce AI features cleanly, such as triage queues, labeling workflows, role-specific review views, and admin controls for thresholds and exceptions. A practical approach is to add AI as a modular layer rather than rebuild the entire product. This layer can include separate inference services, model management, overlays or heatmaps, and review workflows such as triage, review, and disagreement resolution. This helps product companies release AI features faster and iterate on them without destabilizing the core platform.
Make “human-in-the-loop” explicit with safe UX patterns and auditability
AI adoption stalls when pathologists do not trust outputs or cannot explain them. Design your AI pathology software so the human decision is the product’s center: show heatmaps and regions together with confidence scores or confidence levels, provide “why flagged” context, and support easy overrides (confirm, reject, defer). Every AI output should be traceable: model version, data version, thresholds used, and the final human action. This turns your AI from a black box into a reviewable decision-support tool, and it is also the foundation for audits, clinical validation packages, and future regulated pathways.
Design for variability and drift from day one
“It worked in the lab” often fails in production because stain protocols, tissue prep, scanners, compression, and focus artifacts vary across sites. Best practice is to treat variability as a first-class requirement: validate on multi-site data, track performance by scanner, stain, or subtype, and deploy drift monitoring (e.g., slide quality signals, embedding distribution shifts, and error rates by cohort). When confidence drops or drift is detected, route cases to a “human-first” path and trigger an active-learning loop to capture hard cases, label them, and retrain with controlled releases instead of silent model degradation.
Build evaluation like a product feature
AI evaluation should be structured so that every model release is tested the same way and the results can be compared over time. A practical approach is to build a versioned, automated evaluation pipeline with fixed dataset splits, ground-truth provenance, cohort-level metrics, and regression checks for every release. Track not only overall performance metrics, but also failure patterns such as false positives by tissue type, sensitivity by tumor subtype, and performance under slide quality issues. These findings should guide model evolution: they help product teams identify weak spots, refine thresholds, prioritize new data collection and labeling, and target retraining where it matters most.
Optimize WSI pipelines early to avoid cloud-cost surprises
Whole-slide images are very large, so costs rise quickly when systems process full slides unnecessarily, store too much derived data, or keep all image data in high-performance storage. A more efficient approach is to split slides into smaller image patches only when needed for analysis, cache frequently viewed zoom levels or image areas so they do not have to be regenerated or reloaded every time, define how long raw slides and derived data should be retained, and separate frequently accessed data from lower-cost archival storage. For AI inference, product teams should group image patches into efficient processing batches, scale GPU resources up or down based on workload, and track cost per processed slide as an operating metric. If customers need cloud, on-prem, or hybrid deployment, the inference service should be built to run across environments without major rework, for example by packaging it in containers and designing it to work with GPU-enabled infrastructure. This helps product teams control storage and compute costs without turning each deployment into a custom engineering project.
Costs of Developing AI Products for Digital Pathology
The cost of developing an AI product for digital pathology typically ranges from $150,000 to $1,200,000+ and depends mainly on the solution scope and intended use.
Building a research MVP will typically start at $150,000. An assistive AI add-on to an existing pathology platform will normally fall in the $300,000+ range. Building an assistive AI-first pathology product from scratch may require budgets from $900,000. A regulated diagnostic AI module will usually start at $1,200,000+.
Key development cost drivers include:
|
|
Data readiness and labeling: availability and quality of ground truth, dataset preparation effort, labeling complexity, and dataset versioning. |
|
|
WSI pipeline and viewer UX: ingestion, de-identification, tiling logic, caching, multi-format support, heatmaps, overlays, and review workflow screens. |
|
|
Deployment model complexity: cloud, on-prem, or hybrid architecture; GPU-aware deployment; storage architecture; and environment-specific security requirements. |
|
|
Integrations: LIS/LIMS, PACS/DICOM (if needed), SSO, audit exports, research systems and partner APIs. |
|
|
Model scope and robustness requirements: triage vs. segmentation vs. biomarker quantification, and the engineering effort needed to support target performance across scanners, stains, and workflows. |
|
|
MLOps and lifecycle controls: model registry, CI/CD for models, monitoring and alerts, rollback, explainability, versioning, and release controls. |
|
|
Regulated workflow requirements (if applicable): audit trails, role-based access, traceability, validation support features, and documentation support. |
Why ScienceSoft
- In healthcare IT since 2005, with experience building medical imaging AI products.
- 750+ IT experts on board, including AI consultants, data scientists, solution architects, software engineers, QA, DevOps, and compliance specialists.
- Deep knowledge of pathology interoperability, including DICOM whole-slide microscopy for the image layer and HL7 v2/v3 and FHIR for workflow and report exchange.
- ISO 13485, ISO 9001, and ISO 27001 certifications.
- Principal architects with experience in designing scalable healthcare product architectures, secure data exchange, and complex interoperability scenarios.
- Experience supporting compliance with HIPAA/HITECH, GDPR, PDPL, FDA QMSR/21 CFR Part 820, 21 CFR Part 11.
- Knowledge of CLIA/CAP expectations for digital pathology validation and operational readiness.
- Experience working with MDR/IVDR requirements based on intended purpose and target market, with awareness of MDR/IVDR–AI Act interplay for AI-enabled products.
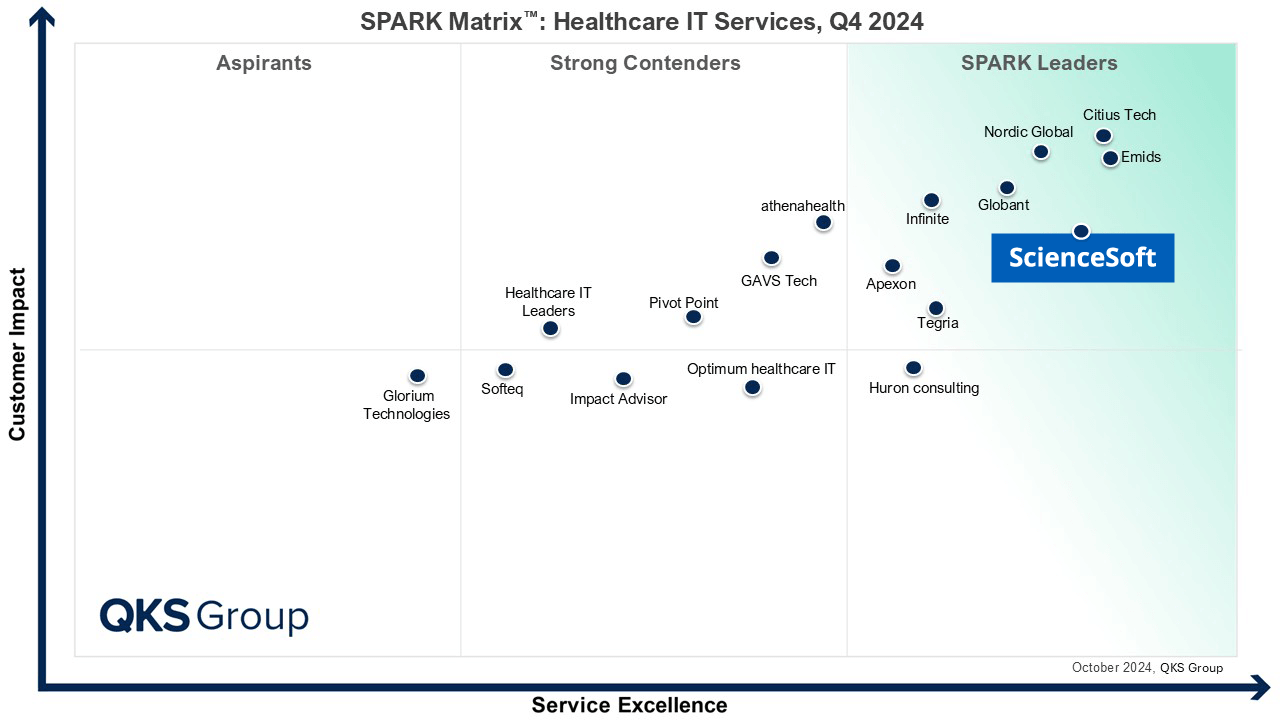
Featured among Healthcare IT Services Leaders in the 2022 and 2024 SPARK Matrix
Recognized for Healthcare Technology Leadership by Frost & Sullivan in 2023 and 2025
Named among America’s Fastest-Growing Companies by Financial Times, 5 years in a row

Top Healthcare IT Developer and Advisor by Black Book™ survey 2023
Four-time finalist across HTN Awards programs

Named to The Healthcare Technology Report’s Top 25 Healthcare Software Companies of 2025

HIMSS Gold member advancing digital healthcare
ISO 13485-certified quality management system
ISO 27001-certified security management system
FAQs
What is the fastest AI feature to launch first?
Worklist triage (priority scoring) and simple cancer-detection heatmaps are usually the fastest because they create immediate productivity value while requiring the least UI and reporting redesign. They also fit naturally into existing review workflows, so they can be adopted without changing how cases are signed out.
How long does it typically take to launch an AI feature in a digital pathology product?
Depending on the scope, initial development and integration of a narrow AI module for research use, for example, a slide quality check, tissue detection, region preselection, or simple heatmap generation, can take about 4–8 months. More advanced or clinical-use capabilities, such as biomarker scoring, case prioritization tied to diagnostic workflows, or AI functions intended for regulated use, usually take significantly longer due to broader workflow design, analytical and clinical validation, and, where applicable, regulatory preparation. As a result, time to market is typically longer than initial development time.
How much labeled data do we need to start?
You need enough labeled slides to represent the variability you will see in production (scanner, stain, tissue prep, and site differences), otherwise accuracy will drop when the model leaves the “training lab.” Many teams start with a narrow use case and a curated dataset, then grow coverage using active learning so labeling effort goes into the most informative new cases.
How do you handle variability across scanners and stains?
We plan for variability up front because it is the most common reason pathology models fail after rollout. That means validating on data from multiple sites and scanners, tracking metrics by scanner and stain, monitoring drift in production, and routing low-confidence cases to manual-first review so performance issues do not silently affect outcomes.
What about scanner format compatibility?
This needs early planning because WSI formats vary by vendor and can affect ingestion, tiling speed, storage cost, and viewer performance. Using proven WSI back ends (e.g., OpenSlide/libvips and tile caching) reduces the risk of format lock when you add new scanner types later.
Research vs. regulated diagnostics: what changes in delivery and cost?
Research projects can move faster because you can iterate more freely and focus on usefulness for discovery and quantification. Regulated diagnostics costs more because AI must be treated as part of a medical device workflow: stricter documentation, traceability, controlled updates, cybersecurity expectations, and deeper validation are required to support regulatory submissions and safe clinical use.
Cloud vs. on-prem inference: what is typical and why?
Cloud inference is often preferred in earlier product stages and initial rollouts because it accelerates delivery and simplifies GPU scaling for computationally intensive whole-slide image processing. On-prem or hybrid inference is more typical when customers have data residency requirements, strict IT policies, or infrastructure preferences that require local deployment. For product vendors, these setups are usually more complex and costly to support because deployment, monitoring, and upgrades must function reliably across heterogeneous customer environments.




